NWAVE Tutorial 1: Regression with Spiking Neural Networks
Tutorial by Giuseppe Gentile and Marco Rasetto
Overview
This tutorial will introduce you on using the NWAVE SDK for training Spiking Neural Networks (SNNs) on a simple task. We'll train a network to learn temporal functions (linear and square-root) and highlight key differences between NWAVE and the popular snnTorch library.
About NWAVE:
NWAVE is designed for training SNNs with hardware deployment in mind. It provides:
- Hardware-ready models: Train SNNs that can be directly deployed on Neuronova's neuromorphic chips
- Easy migration: Port existing networks from other frameworks with minimal code changes.
- PyTorch compatibility: Standard
[B, T, N]data format familiar to deep learning practitioners - Time series parallelization: Train LIF models via ComPaSSo, a GPU acceleration method to speed execution and training of SNNs
Reference: This tutorial follows a similar approach to the snnTorch regression tutorial to facilitate direct comparison.
What You'll Learn
- How to create and train LIF(Leaky Integrate and Fire) SNNs using NWAVE
- Key differences in data format between NWAVE and snnTorch
- Training and evaluating SNNs for regression tasks
- Performance monitoring and visualization
1. Setup and Imports
import torch
import torch.nn as nn
from torch.utils.data import DataLoader
import matplotlib.pyplot as plt
import numpy as np
import random
import tqdm
# NWAVE imports
from nwavesdk import LIFSynapse, LIFLayer, prepare_net
Set Random Seeds for Reproducibility
torch.manual_seed(42)
random.seed(42)
np.random.seed(42)
2. Dataset Generation
We create a simple regression dataset where the task is to learn temporal functions. The dataset generates linear trajectories from 0 to a random endpoint, and the target can be either:
- Linear: Direct copy of the input
- Square-root: Non-linear transformation of the input
This is identical to the snnTorch tutorial dataset for fair comparison.
class RegressionDataset(torch.utils.data.Dataset):
"""Simple regression dataset for temporal functions."""
def __init__(self, timesteps, num_samples, mode):
"""Generate dataset with linear or square-root relationship.
Args:
timesteps: Number of time steps in each sequence
num_samples: Number of samples to generate
mode: 'linear' or 'sqrt' for the target function type
"""
self.num_samples = num_samples
feature_lst = []
# Generate linear functions one by one
for idx in range(num_samples):
end = float(torch.rand(1)) # Random final point
lin_vec = torch.linspace(start=0.0, end=end, steps=timesteps)
feature = lin_vec.view(timesteps, 1)
feature_lst.append(feature)
self.features = torch.stack(feature_lst, dim=1) # [T, B, N]
# Generate labels based on mode
if mode == "linear":
self.labels = self.features * 1
elif mode == "sqrt":
slope = float(torch.rand(1))
self.labels = torch.sqrt(self.features * slope)
else:
raise NotImplementedError("mode must be 'linear' or 'sqrt'")
def __len__(self):
return self.num_samples
def __getitem__(self, idx):
return self.features[:, idx, :], self.labels[:, idx, :]
Dataset Parameters and Visualization
sim_dt = 1e-3 # The dt in seconds for the simulation
num_steps = 50 # In the dt of the simulation, 50ms
num_samples = 1024
mode = "sqrt" # Options: 'linear' or 'sqrt'
# Generate dataset
dataset = RegressionDataset(timesteps=num_steps, num_samples=num_samples, mode=mode)
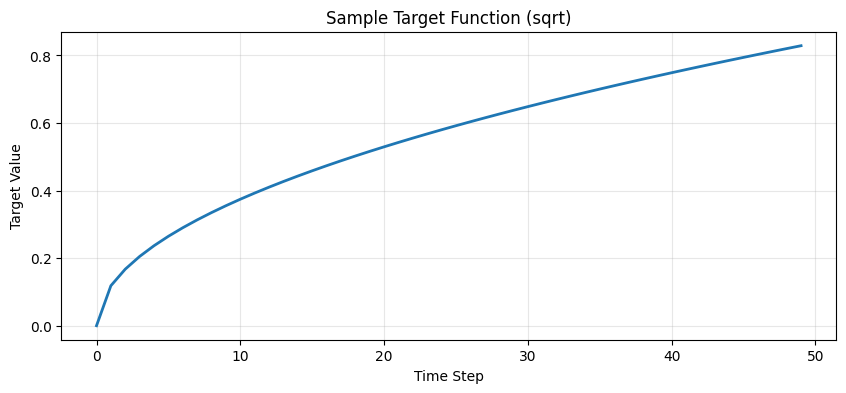
# Visualize a sample target function
sample = dataset.labels[:, 0, 0]
plt.figure(figsize=(10, 4))
plt.plot(sample, linewidth=2)
plt.title(f"Sample Target Function ({mode})")
plt.xlabel("Time Step")
plt.ylabel("Target Value")
plt.grid(True, alpha=0.3)
plt.show()
3. Key Difference: Data Format
NWAVE vs snnTorch Data Representation
This is one of the most important differences between the two frameworks:
| Framework | Data Format | Description |
|---|---|---|
| snnTorch | [T, B, N] |
Time-first format, aligns with SNN temporal processing |
| NWAVE | [B, T, N] |
Batch-first format, consistent with standard PyTorch conventions |
Where:
- B: Batch size (number of samples)
- T: Time steps (sequence length)
- N: Features (dimensions per time step)
Why this matters:
- snnTorch uses
[T, B, N]because it processes spikes temporally, iterating over time steps first - NWAVE uses
[B, T, N]to maintain consistency with standard deep learning frameworks (PyTorch RNNs, Transformers, etc.) and to facilitate easier integration with existing data pipelines
Hardware deployment advantage: The [B, T, N] format in NWAVE makes it easier to:
- Port models from standard PyTorch implementations
- Work with existing data processing pipelines
- Prepare batches for efficient hardware deployment
In practice: When using data prepared for snnTorch with NWAVE, you need to permute dimensions from [T, B, N] to [B, T, N].
print(f"Original dataset shape (snnTorch format): {dataset.labels.shape}")
print(f"Format: [Time={dataset.labels.shape[0]}, Batch={dataset.labels.shape[1]}, Features={dataset.labels.shape[2]}]")
Original dataset shape (snnTorch format): torch.Size([50, 1024, 1])
Format: [Time=50, Batch=1024, Features=1]
Create DataLoader
The DataLoader will automatically handle the permutation when we iterate through batches.
batch_size = 32
dataloader = torch.utils.data.DataLoader(
dataset=dataset, batch_size=batch_size, drop_last=True, shuffle=True
)
# Check the shape after DataLoader
sample_batch = next(iter(dataloader))
print(f"\nDataLoader output shape (NWAVE format): {sample_batch[0].shape}")
print(f"Format: [Batch={sample_batch[0].shape[0]}, Time={sample_batch[0].shape[1]}, Features={sample_batch[0].shape[2]}]")
DataLoader output shape (NWAVE format): torch.Size([32, 50, 1])
Format: [Batch=32, Time=50, Features=1]
4. Building the SNN Model
Network Architecture
We'll create a simple two-layer feedforward SNN using Leaky Integrate-and-Fire (LIF) neurons.
Model Parameters
input_size = 1 # Single-dimensional input
hidden_size = 256 # Hidden layer neurons
output_size = 1 # Single-dimensional output
# For ComPaSSo check ComPaSSo Tutorial
device = "cpu" # NWAVE uses "gpu" if GPU available for GPU acceleration via ComPaSSo
5. Model Definition: NWAVE vs snnTorch
Key Architectural Differences
| Aspect | snnTorch | NWAVE |
|---|---|---|
| Layer Structure | Separate nn.Linear + snn.Leaky |
Combined LIFSynapse + LIFLayer |
| Time Loop | Manual loop over time steps | Manual loop over time steps |
| State Management | Returns (spikes, membrane) | Returns (spikes, membrane) |
| Bias Learning | Standard PyTorch | Explicit bias_learn parameter |
| Neuron Parameters | Abstract (beta decay) | Physical (tau, dt, threshold) |
| Reset Mechanism | Default subtract | Explicit reset_mechanism parameter |
| Data Format | [T, B, N] input |
[B, T, N] input |
| Hardware Deployment | Simulation-focused | Hardware-ready models |
Why we choose Physical Taus
NWAVE uses physical neuron parameters (time constants in seconds, simulation dt in seconds) instead of abstract parameters. This is critical for Hardware mapping as our analog solutions compute information in realtime on temporal scales in the order milliseconds, making hardware deployment more natural.
NWAVE Model Implementation
Understanding LIF Neuron Parameters
Before building the model, let's understand how the physical parameters in LIFLayer work:
The LIF Neuron Model
The Leaky Integrate-and-Fire neuron follows these dynamics:
# 1. Leaky Integration (membrane potential decay + input accumulation)
tau * dV/dt = -V + R*I(t)
# Discrete approximation:
V[t+1] = V[t] * exp(-dt/tau) + I[t]
Where:
- V: Membrane potential (voltage)
- tau (τ): Membrane time constant - how fast the neuron "forgets" past inputs
- dt: Simulation time step
- I(t): Input current at time t
- R: Membrane resistance (absorbed into input current in practice)
# 2. Spike Generation
if V[t] >= threshold:
spike = 1
else:
spike = 0
# 3. Reset Mechanism
if spike == 1:
V[t] = V[t] - threshold # "subtraction" mode
# OR
V[t] = 0 # "zero" mode
Parameter Interpretation
| Parameter | Typical Range | Biological Meaning | Effect on Network |
|---|---|---|---|
| tau | 5-30 ms | Membrane time constant | Larger tau → slower response, more temporal integration |
| dt | 0.1-1 ms | Simulation timestep | Smaller dt → more accurate but slower simulation |
| threshold | 0.1-1.0 | Spike threshold | Higher threshold → fewer spikes, harder to activate |
| reset_mechanism | "subtraction" | How membrane resets | "subtraction" preserves excess charge, "zero" discards it |
Example Parameter Choices in This Tutorial
Hidden Layer (neu1):
taus = 16e-3 # 16ms - medium integration window
thresholds = 0.2 # Low threshold - easier to spike
dt = 1e-3 # 1ms timestep
reset = "subtraction"
- Why 16ms tau? Good balance between responsiveness and temporal integration for the 50ms dataset
- Why 0.2 threshold? Low enough to allow frequent spiking for learning
- Why subtraction reset? Preserves information about "how much" threshold was exceeded
Output Layer (neu2):
taus = 1e-3 # 1ms - very fast (almost no memory)
thresholds = 10 # Very high - essentially won't spike!
dt = 1e-3
reset = "subtraction"
- Why 1ms tau? Minimal temporal integration - acts like a "readout" layer
- Why threshold=10? So high that it never spikes - we read the membrane potential, not spikes
- Why this design? For regression, we want continuous outputs, so we use membrane voltage as the output signal
class NWaveRegressionNet(nn.Module):
"""NWAVE SNN for regression task.
Architecture:
- Layer 1: 1 -> 256 LIF neurons (hidden layer)
- Layer 2: 256 -> 1 LIF neuron (output layer)
"""
def __init__(self):
super().__init__()
# Layer 1: Input -> Hidden
self.syn1 = LIFSynapse(input_size, hidden_size, use_bias=True, bias_learn=True)
self.neu1 = LIFLayer(
hidden_size,
taus=16e-3, # Time constant (16ms)
thresholds=0.2, # Spike threshold
reset_mechanism="subtraction", # Subtract threshold on spike
dt=sim_dt # Time step (1ms)
)
# Layer 2: Hidden -> Output
self.syn2 = LIFSynapse(hidden_size, output_size, use_bias=True, bias_learn=True)
self.neu2 = LIFLayer(
output_size,
taus=1e-3, # Shorter time constant for output
thresholds=10, # High threshold (membrane readout)
reset_mechanism="subtraction",
dt=sim_dt
)
def forward(self, x):
"""Forward pass through the network.
Args:
x: Input tensor of shape [B, T, N]
Returns:
spk_out: Output spikes [B, T, 1]
mem_out: Output membrane potentials [B, T, 1]
spk_hidden: Hidden layer spikes [B, T, hidden_size]
"""
B, T, _ = x.shape
mem_trace = []
spk_trace = []
spk_trace_hidden = []
# Prepare network: Initialize hidden states (membrane potentials)
# This MUST be called before the forward pass to reset internal states
# Unlike snnTorch where you pass mem as input, NWAVE manages state internally
prepare_net(self)
# Time loop - process each time step
for t in range(T):
# Layer 1
cur1 = self.syn1(x[:, t, :]) # Synaptic current
spk1, mem1 = self.neu1(cur1) # Neuron dynamics
spk_trace_hidden.append(spk1)
# Layer 2
cur2 = self.syn2(spk1)
spk2, mem2 = self.neu2(cur2)
mem_trace.append(mem2)
spk_trace.append(spk2)
# Stack temporal outputs: [B, T, N]
mem_trace = torch.stack(mem_trace, dim=1)
spk_trace = torch.stack(spk_trace, dim=1)
spk_trace_hidden = torch.stack(spk_trace_hidden, dim=1)
return spk_trace, mem_trace, spk_trace_hidden
# Instantiate the model
model = NWaveRegressionNet().to(device)
print("\n=== Model Summary ===")
print(f"Total parameters: {sum(p.numel() for p in model.parameters()):,}")
print(f"Trainable parameters: {sum(p.numel() for p in model.parameters() if p.requires_grad):,}")
=== Model Summary ===
Total parameters: 769
Trainable parameters: 769
snnTorch Equivalent (for reference)
# This is how the same network would look in snnTorch:
import snntorch as snn
class SNNTorchRegressionNet(nn.Module):
def __init__(self):
super().__init__()
# Layer 1
self.fc1 = nn.Linear(input_size, hidden_size)
self.lif1 = snn.Leaky(beta=0.9, init_hidden=True)
# Layer 2
self.fc2 = nn.Linear(hidden_size, output_size)
self.lif2 = snn.Leaky(beta=0.9, init_hidden=True)
def forward(self, x):
# x shape: [T, B, N] for snnTorch
# IMPORTANT: Manual state initialization in snnTorch
mem1 = self.lif1.init_leaky()
mem2 = self.lif2.init_leaky()
spk_rec = []
mem_rec = []
for step in range(x.size(0)): # Iterate over time
cur1 = self.fc1(x[step])
spk1, mem1 = self.lif1(cur1, mem1) # Pass mem as input AND output!
cur2 = self.fc2(spk1)
spk2, mem2 = self.lif2(cur2, mem2) # Manual state tracking
spk_rec.append(spk2)
mem_rec.append(mem2)
return torch.stack(spk_rec), torch.stack(mem_rec)
Critical Differences:
-
State Management ⭐ Most Important Difference
- snnTorch: Manual state management - you must pass
memas both input and outputpython mem = lif.init_leaky() # Initialize spk, mem = lif(current, mem) # Pass mem explicitly - NWAVE: Automatic internal state management - use
prepare_net()oncepython prepare_net(model) # Initialize all layers spk, mem = lif(current) # No mem argument needed! - Why it matters: NWAVE's approach is cleaner, less error-prone, and mirrors hardware behavior
- snnTorch: Manual state management - you must pass
-
Layer Architecture
- NWAVE: Separates synaptic connections (
LIFSynapse) from neuron dynamics (LIFLayer) - snnTorch: Combines both in standard
nn.Linearfollowed by spiking neuron layers - Why it matters: NWAVE's separation mirrors hardware architecture where synapses and neurons are distinct components
- NWAVE: Separates synaptic connections (
-
Neuron Parameters
- NWAVE:
taus=16e-3(16ms),dt=1e-3(1ms), Physical values! - snnTorch:
beta=0.9- Abstract decay parameter - Conversion:
beta = exp(-dt/tau), sotau ≈ -dt/log(beta) ≈ 9.5msfor beta=0.9, dt=1ms - Why it matters: Physical parameters in NWAVE map directly to chip configurations
- NWAVE:
-
Data Format
- NWAVE:
[B, T, N]- batch first (standard PyTorch) - snnTorch:
[T, B, N]- time first (SNN-specific) - Why it matters: Easier to port existing PyTorch models to NWAVE
- NWAVE:
-
Function Calls Per Timestep
- NWAVE:
prepare_net()once, then simplelayer(input)calls - snnTorch: Must track and pass
memvariables for every neuron layer at every timestep - Why it matters: NWAVE reduces boilerplate code and potential bugs
- NWAVE:
Migration from snnTorch to NWAVE - Quick Guide:
| snnTorch Code | NWAVE Equivalent |
|---|---|
nn.Linear(in, out) + snn.Leaky(beta) |
LIFSynapse(in, out) + LIFLayer(out, taus, thresholds, dt) |
mem = lif.init_leaky() |
prepare_net(model) (once before forward) |
spk, mem = lif(cur, mem) |
spk, mem = lif(cur) (no mem argument!) |
x.shape = [T, B, N] |
x.shape = [B, T, N] (permute dimensions) |
beta = 0.9 |
tau = -dt/log(beta) ≈ 9.5ms for dt=1ms |
Understanding NWAVE SDK Functions
NWAVE provides specialized functions and layers that handle SNN simulation and state management differently from snnTorch. Here are the key components:
1. LIFSynapse(nb_inputs, nb_outputs, use_bias, bias_learn)
Purpose: Represents synaptic connections (weights) between neuron layers.
Parameters:
nb_inputs: Number of input neuronsnb_outputs: Number of output neuronsuse_bias: Whether to include bias termsbias_learn: Whether bias is trainable (separate from weight learning)
Key Features:
- Implements weighted connections:
output = input @ weights + bias - Supports quantization for hardware deployment (
quantization_bits) - Xavier initialization for stable training
- Can build quantized weights via
build_Q()method
Hardware mapping: Synaptic weights map directly to crossbar arrays in neuromorphic chips.
2. LIFLayer(n_neurons, taus, thresholds, reset_mechanism, dt, ...)
Purpose: Implements Leaky Integrate-and-Fire neuron dynamics with internal state management.
Parameters:
n_neurons: Number of neurons in the layertaus: Membrane time constant(s) in seconds (e.g., 16e-3 = 16ms)thresholds: Spike threshold value(s)reset_mechanism: How membrane resets after spike ("subtraction", "zero", or "none")dt: Simulation time step in seconds (e.g., 1e-3 = 1ms)spike_grad: Surrogate gradient function for backpropagationlayer_topology: "FF" (feedforward) or "RC" (recurrent)
Key Features:
- Internal state management: Membrane potential is stored inside the layer (not passed as input!)
- Reset mechanism options:
"subtraction":mem = mem - threshold(most common)"zero":mem = 0(hard reset)"none": No reset (membrane keeps accumulating)
Mathematical model:
# Membrane dynamics (leaky integration):
mem_new = mem_old * exp(-dt/tau) + input_current
# Spike generation:
spike = 1 if mem_new >= threshold else 0
# Reset after spike:
mem_new = reset_function(mem_new, spike)
Hardware mapping:
tauandthresholddirectly configure analog neuron circuitsdtdetermines the discrete time step for digital event processing
3. prepare_net(model)
Purpose: Initializes/resets the network state before running a forward pass.
Why this is needed:
- SNNs are stateful - neurons maintain membrane potentials across time steps
- Must reset states between different input sequences (different batches)
- Ensures clean state initialization for each forward pass
Key difference from snnTorch:
# snnTorch: Manual state management
mem1 = lif1.init_leaky() # Initialize
for t in range(timesteps):
spk, mem1 = lif1(cur, mem1) # Pass mem as input/output
# NWAVE: Automatic state management
prepare_net(model) # Initialize once
for t in range(timesteps):
spk, mem = lif_layer(cur) # State managed internally!
Benefits:
- Cleaner code - no need to track
memvariables manually - Less error-prone - harder to forget state initialization
- Better hardware alignment - mirrors how neuromorphic chips manage state
This internal state management is why you don't pass mem as an argument in NWAVE!
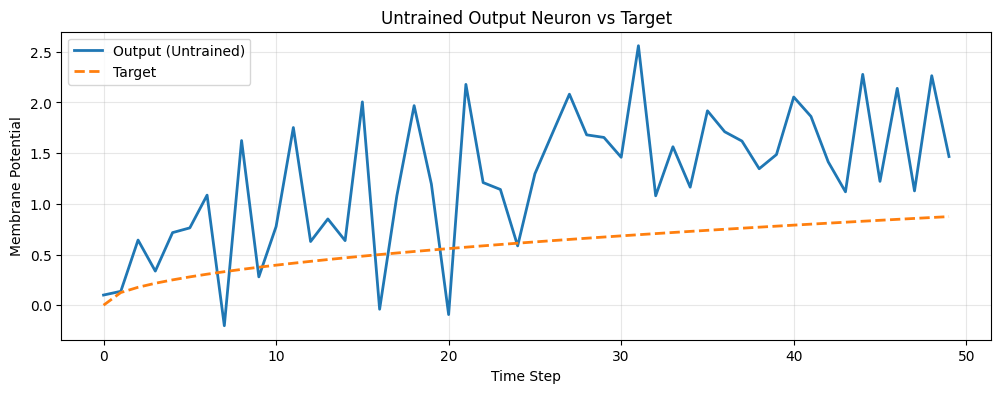
6. Untrained Network Behavior
Let's observe how the network behaves before training. The output should be random and not match the target.
Key Point: When to Call prepare_net()
The prepare_net() function is called inside the forward method in this tutorial because:
- State Reset Between Sequences: Each batch is a different sequence, so we reset membrane potentials to zero
- Metric Collection: We enable spike rate tracking for monitoring network activity
- Clean Initialization: Ensures all neurons start from a known state
Important: In production or when processing multiple batches sequentially, you might want to:
- Call
prepare_net()once before the time loop (not inside) - Only reset between different input sequences, not between timesteps
- Preserve state across timesteps within a sequence
Example patterns:
# Pattern 1: Reset per batch (as in this tutorial)
def forward(self, x):
prepare_net(self, collect_metrics=True) # Reset for each batch
for t in range(T):
spk, mem = self.neu(self.syn(x[:, t]))
return spk, mem
# Pattern 2: Reset only at sequence start (more efficient)
prepare_net(model, collect_metrics=True) # Call once before training
for batch in dataloader:
output = model(batch) # State persists across batches
# Pattern 3: Manual reset control
model.neu1.reset() # Reset specific layer only
For this tutorial, we use Pattern 1 for clarity and to ensure clean state per forward pass.
# Get a batch of data
train_batch = iter(dataloader)
feature, label = next(train_batch)
# Run forward pass without gradients
with torch.no_grad():
feature = feature.to(device)
label = label.to(device)
spk, mem, spk_h = model(feature)
# Visualize first sample
plt.figure(figsize=(12, 4))
plt.plot(mem[0, :, 0].cpu(), label="Output (Untrained)", linewidth=2)
plt.plot(label[0, :, 0].cpu(), "--", label="Target", linewidth=2)
plt.title("Untrained Output Neuron vs Target")
plt.xlabel("Time Step")
plt.ylabel("Membrane Potential")
plt.legend(loc="best")
plt.grid(True, alpha=0.3)
plt.show()
# Check initial spike rate
print(f"\nOutput layer spike rate: {spk.mean().item():.4f}")
print(f"Hidden layer spike rate: {spk_h.mean().item():.4f}")
Output layer spike rate: 0.0000
Hidden layer spike rate: 0.4659
7. Training the Network
Training Configuration
We'll train the network to minimize the Mean Squared Error (MSE) between the output membrane potential and the target function.
# Training hyperparameters
num_epochs = 150
learning_rate = 1e-2
# Optimizer and loss function
optimizer = torch.optim.AdamW(params=model.parameters(), lr=learning_rate)
loss_function = torch.nn.MSELoss()
loss_hist = [] # Track loss over time
# Training loop
model.train()
with tqdm.trange(num_epochs) as pbar:
for epoch in pbar:
epoch_loss = []
for feature, label in dataloader:
feature = feature.to(device)
label = label.to(device)
# Forward pass
spk_out, mem_out, spk_hidden = model(feature)
# Compute loss on membrane potential
loss = loss_function(mem_out, label)
# Backward pass
optimizer.zero_grad()
loss.backward()
optimizer.step()
# Record loss
loss_hist.append(loss.item())
epoch_loss.append(loss.item())
# Update progress bar
avg_loss = sum(epoch_loss) / len(epoch_loss)
pbar.set_postfix(loss="%.3e" % avg_loss)
print("\nTraining completed!")
100%|██████████| 150/150 [00:37<00:00, 4.03it/s, loss=6.902e-04]
Training completed!
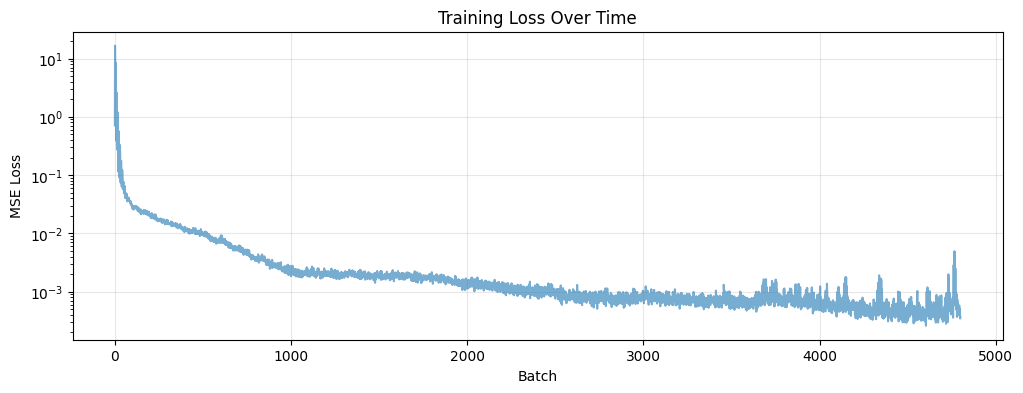
Training Loss Visualization
plt.figure(figsize=(12, 4))
plt.plot(loss_hist, alpha=0.6)
plt.title("Training Loss Over Time")
plt.xlabel("Batch")
plt.ylabel("MSE Loss")
plt.yscale('log')
plt.grid(True, alpha=0.3)
plt.show()
8. Evaluation and Results
Let's evaluate the trained network and visualize its performance.
# Switch to evaluation mode
model.eval()
# Get test batch
test_batch = iter(dataloader)
feature, label = next(test_batch)
# Evaluate without gradients
with torch.no_grad():
feature = feature.to(device)
label = label.to(device)
spk, mem, spk_h = model(feature)
# Move to CPU for plotting
mem = mem.cpu()
label = label.cpu()
spk = spk.cpu()
spk_h = spk_h.cpu()
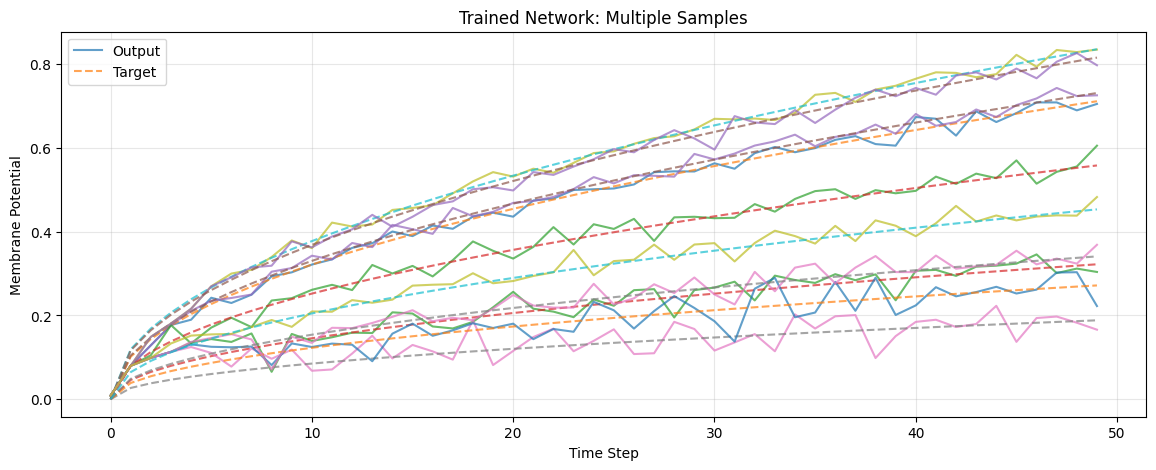
Visualize Multiple Predictions
plt.figure(figsize=(14, 5))
plt.title("Trained Network: Multiple Samples")
plt.xlabel("Time Step")
plt.ylabel("Membrane Potential")
# Plot multiple samples from the batch
num_samples_to_plot = min(10, batch_size)
for i in range(num_samples_to_plot):
out_trace = mem[i, :, 0]
target_trace = label[i, :, 0]
# Only add labels for the first sample to avoid clutter
plt.plot(out_trace, alpha=0.7, label="Output" if i == 0 else None)
plt.plot(target_trace, "--", alpha=0.7, label="Target" if i == 0 else None)
plt.legend(loc="best")
plt.grid(True, alpha=0.3)
plt.show()
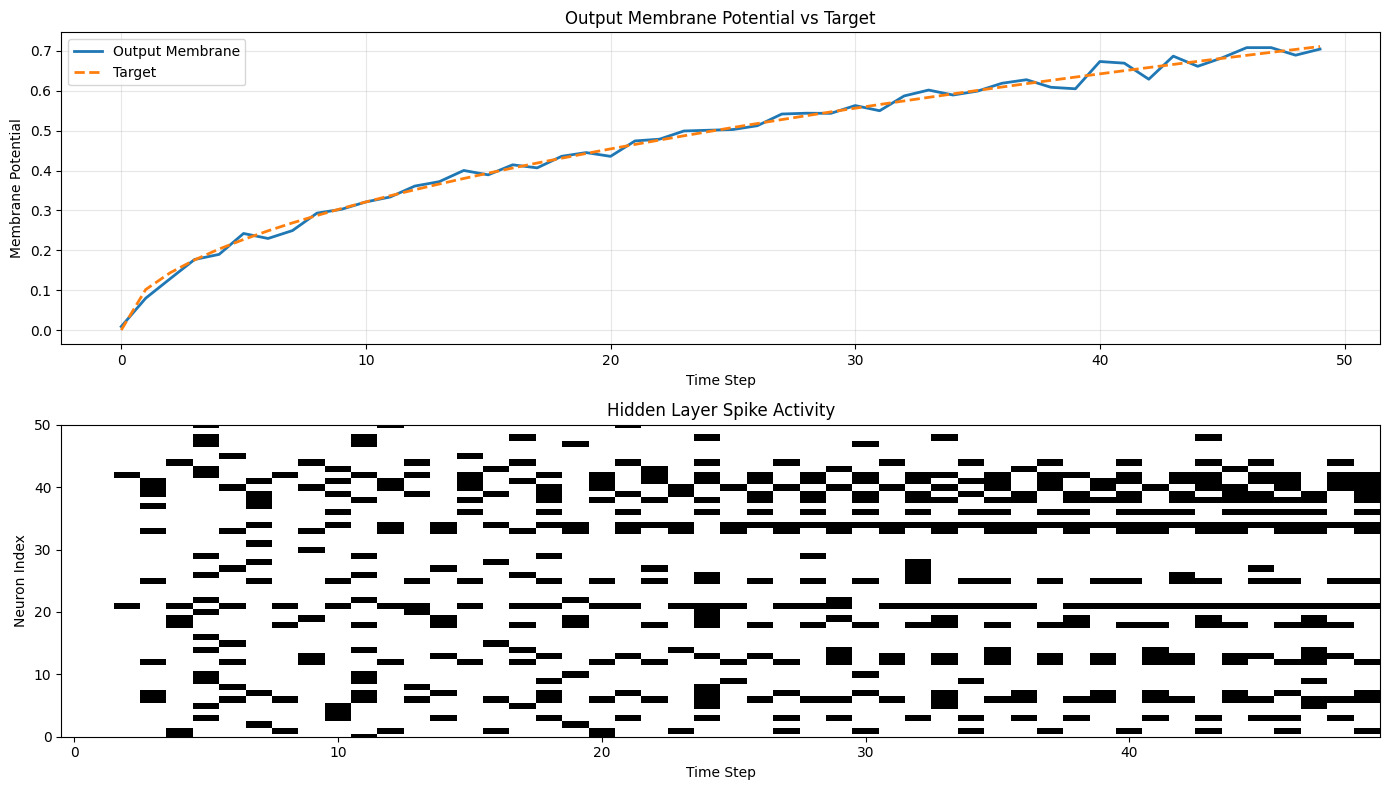
Detailed Single Sample Analysis
# Detailed view of a single sample
sample_idx = 0
fig, axes = plt.subplots(2, 1, figsize=(14, 8))
# Plot 1: Membrane potential
axes[0].plot(mem[sample_idx, :, 0], label="Output Membrane", linewidth=2)
axes[0].plot(label[sample_idx, :, 0], "--", label="Target", linewidth=2)
axes[0].set_title("Output Membrane Potential vs Target")
axes[0].set_xlabel("Time Step")
axes[0].set_ylabel("Membrane Potential")
axes[0].legend()
axes[0].grid(True, alpha=0.3)
# Plot 2: Hidden layer activity
axes[1].imshow(spk_h[sample_idx, :, :].T, aspect='auto', cmap='binary', interpolation='nearest')
axes[1].set_title("Hidden Layer Spike Activity")
axes[1].set_xlabel("Time Step")
axes[1].set_ylabel("Neuron Index")
axes[1].set_ylim([0, min(50, hidden_size)]) # Show first 50 neurons
plt.tight_layout()
plt.show()
print(f"\nFinal spike rate (output): {spk.mean().item():.4f}")
print(f"Final spike rate (hidden): {spk_h.mean().item():.4f}")
Final spike rate (output): 0.0000
Final spike rate (hidden): 0.2043
Performance Metrics
# Compute various metrics
mse = torch.nn.functional.mse_loss(mem[:, :, 0], label[:, :, 0])
mae = torch.nn.functional.l1_loss(mem[:, :, 0], label[:, :, 0])
# R² score
ss_res = torch.sum((label[:, :, 0] - mem[:, :, 0]) ** 2)
ss_tot = torch.sum((label[:, :, 0] - label[:, :, 0].mean()) ** 2)
r2_score = 1 - (ss_res / ss_tot)
print("\n=== Performance Metrics ===")
print(f"Mean Squared Error (MSE): {mse.item():.6f}")
print(f"Mean Absolute Error (MAE): {mae.item():.6f}")
print(f"R² Score: {r2_score.item():.6f}")
# Network activity metrics
print("\n=== Network Activity ===")
print(f"Output layer spike rate: {spk.mean().item():.4f}")
print(f"Hidden layer spike rate: {spk_h.mean().item():.4f}")
print(f"Active neurons (>0.1% spike rate): {(spk_h.mean(dim=[0,1]) > 0.001).sum().item()} / {hidden_size}")
=== Performance Metrics ===
Mean Squared Error (MSE): 0.000516
Mean Absolute Error (MAE): 0.016603
R² Score: 0.989620
=== Network Activity ===
Output layer spike rate: 0.0000
Hidden layer spike rate: 0.2043
Active neurons (>0.1% spike rate): 244 / 256
9. Summary: NWAVE vs snnTorch
Quick Comparison Table
| Feature | NWAVE | snnTorch |
|---|---|---|
| Primary Purpose | Hardware deployment on Neuronova chips | SNN research and simulation |
| State Management | Automatic (internal to layers) | Manual (pass mem as argument) |
| Initialization | prepare_net(model) once |
mem = layer.init_leaky() per layer |
| Data Format | [B, T, N] (batch-first) |
[T, B, N] (time-first) |
| Layer Definition | LIFSynapse + LIFLayer |
nn.Linear + snn.Leaky |
| Parameter Style | Physical (tau, dt, thresholds) | Abstract (beta) |
| Bias Control | Explicit bias_learn |
Standard PyTorch |
| Reset Mechanism | Explicit parameter | Fixed per neuron type |
| PyTorch Integration | Standard conventions | SNN-specific conventions |
| Hardware Deployment | ✅ Direct export to neuromorphic chips | ❌ Simulation only |
Migration Guide: snnTorch → NWAVE
Quick reference for porting existing code:
| Aspect | snnTorch | NWAVE |
|---|---|---|
| Imports | import snntorch as snn |
from nwavesdk import LIFSynapse, LIFLayer, prepare_net |
| Synapse | nn.Linear(in, out) |
LIFSynapse(in, out, use_bias=True, bias_learn=True) |
| Neuron | snn.Leaky(beta=0.9) |
LIFLayer(n, taus=10e-3, thresholds=0.5, reset_mechanism="subtraction", dt=1e-3) |
| Beta → Tau | beta = 0.9 |
tau = -dt / log(beta) ≈ 9.5ms (for dt=1ms) |
| Init State | mem = lif.init_leaky() |
prepare_net(model) |
| Forward | spk, mem = lif(cur, mem) |
spk, mem = lif(cur) |
| Data Shape | [T, B, N] |
[B, T, N] (use .permute(1,0,2)) |
Next Steps:
- Try different neuron parameters (tau, threshold) to understand hardware constraints
- Experiment with different network architectures (deeper, recurrent)
- Explore the
layer_topology="RC"option for recurrent connections - Check out Tutorial 2 for classification tasks with spike-based encoding
- Learn about mismatch and quantization for Neuronova hardware deployment
- Experiment with different reset mechanisms ("subtraction", "zero", "none")
Learn More:
- Neuronova Technology - Hardware specifications and capabilities
- NWAVE Documentation - Check your SDK installation for detailed API reference
- Contact Neuronova for hardware deployment support and deployment tools